Articles
AI solution to Levinthal’s paradox
March 2021
DeepMind’s AI program has cracked a fiendishly complex biological problem that had remained unsolved for half a century, and has amazed scientists by demonstrating a capability to predict protein folding with incredible accuracy.
‘Alphafold 2’, an artificial intelligence tool for generating 3D models of proteins based on genomic datasets, was announced in November 2020 and hailed as one of the most ground-breaking scientific technologies of our time, allowing scientists to see how a protein will fold based merely on its primary amino acid sequence without the need for laborious wet techniques. But what are the issues with IP generated from such a useful tool?
Breakthrough
DeepMind is an AI company founded in London in 2010 to focus on machine learning and systems neuroscience with a vision of “making computers think like humans” and was acquired by Google in 2014. DeepMind had widely-publicised success in 2015 with its AlphaGo program, which was the first computer program to defeat a professional human Go player, and has more recently been developing an artificial intelligence tool which predicts protein folding from only a primary amino acid sequence. The latest version of the tool, named ‘AlphaFold 2’, has recently achieved staggering results in the annual protein structure prediction challenge, CASP [1].
The results of the challenge are announced each year at the annual conference which has taken place since 1994, and the team which provides the most accurate structural prediction are pronounced winners. In 2020, the AlphaFold 2 outperformed all other teams by some margin, reaching a score of 92.4 GDT (out of 100) overall across all targets, meaning that its predictions have an average error (RMSD) of only 1.6 Angstroms [2].
Why is this significant?
Scientists are already using the tool to solve protein structures that they have previously struggled with for years, but in a matter of hours. The structure of proteins is typically studied by crystallising the protein and visualising it using X-ray crystallography or more recently cryo-electron microscopy and NMR. However this process is slow, laborious and expensive, and some proteins can be difficult to crystallise such as those embedded in membranes. This difficulty is sometimes referred to as Levinthal’s paradox; Levinthal estimated 10300 possible conformations for a typical protein. Yet in nature, proteins fold spontaneously, some within milliseconds, into a predetermined defined structure.
AlphaFold 2 vastly accelerates the process of understanding how a protein folds and what it looks like in 3D space based only on the primary amino acid sequence.
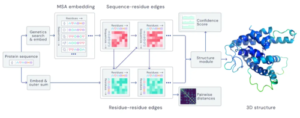
Figure 1: the basis of the AlphaFold 2 system (image credit: https://en.wikipedia.org/w/index.php?curid=65998083)
Visualisation of protein structure has uses in understanding how cellular interactions occur. In particular how proteins bind to other molecules and perform their functions, and what parts of the protein structure are key to this. In essence, how a protein works is thought to be largely determined by its 3D shape.
Having such knowledge easily accessible will advance many fields of life sciences research and likely lead to many new therapies. It will accelerate drug discovery in which candidate drugs are often screened for binding to a particular protein, typically to modify its function in vivo, and provide a therapeutic effect. It will also accelerate the development of useful novel protein design, where, for example, a particular residue which is key to the function can be modified to increase or decrease its activity. It will also help accelerate the identification of structural differences resulting from disease-causing gene variations, helping to arrive at treatments for such genetic diseases more quickly. Understandably, the advancement is being hailed as one of the most significant breakthroughs of the century.
Potential IP issues
Given the significant advantages of AlphaFold 2, undoubtedly many inventions will result from its use. Novel small molecule drugs, novel proteins and their medical or industrial uses are just a few. However, are such inventions patentable, and what potential issues are there with patentability? Also, if an AI creates intellectual property such as an invention, or part of an invention, is it an inventor even though it is not a person? If there is no human inventor then can the invention be patented at all?
Patentability
Leaving aside the inventorship question for now, it is undoubtedly possible for many AI-related inventions to be protected by a patent in Europe. In order to be patentable, it is necessary for an AI-related invention to provide a technical effect, which may be, for example, based on a real-world application of an AI tool, the output from an AI tool, an innovative way of preparing training data for input to an AI system, or even an innovative AI processing hardware arrangement per se. In the biological application sphere, it is conceivable that a new use case for a protein, or a novel protein whose folded structure has been visualised using the AlphaFold 2 tool (or similar such tools) could form the basis for a patentable invention.
However, the use of AlphaFold 2, much like other in silico modelling programs, perhaps raises questions over how inventive step is assessed. In particular, given that a tool now exists to solve the structure of proteins quickly and easily there is certainly less difficulty associated with analysing proteins and identifying useful solutions directly from such structures. It is therefore possible that patent offices will consider solutions resulting from the realisation that a given protein binds to a certain partner such as a receptor or drug more obvious, when the structure clearly shows a known binding region for such partners, and when screening for binding could potentially be done entirely in silico. In other words, when an AI tool is readily available for predicting a protein structure in silico, should the threshold for inventive step be raised relative to the inventive step required had the AI protein folding algorithm not been available?
Advancements in protein folding predictions could also give rise to an increase in “insufficient” reach-through claims – claims that cover potential novel proteins and/or potential uses of the protein that have been identified using the AI tool, but which have not actually been tested and demonstrated to work. For example, this situation could arise if researchers were to use AlphaFold 2 to identify the 3D structure of a novel protein, hypothesise what it could bind to and infer resulting potential uses, and then attempt to claim these uses without further testing. Alternatively, claims may be filed which attempt to cover any binding partner of a given resolved protein structure which could be useful. Such claims were prevalent when cloning and gene sequencing technology first became widespread in the 1980s, allowing scientists to identify genes that had not previously been identified. Many applicants attempted to claim these gene sequences and their hypothetical uses based merely on sequence information [4]. In some cases in the US, applications were filed in which thousands of sequence tags were covered that had no known function. This type of claim is now seen as lacking enablement or support if there is no proper data demonstrating a technical use for the isolated sequence. We may well see a rise in such speculative filings once again, but this time based on proteins rather than genes.
AI Inventorship
It would appear, at least for the moment, that the AlphaFold 2 tool is for aiding the inventive efforts of human scientists rather than for devising inventions autonomously. Accordingly, it seems likely that inventions derived using the AlphaFold 2 tool would still have human inventors. That said, it is perhaps possible that future developments of the AlphaFold system could be more autonomous, for example determining the protein structure and processing this to find the binding partners or particular applications without the expertise of scientists. Where humans merely test the suitability of a shortlist of suggested molecules output by such an AI-tool for their associated applications, identifying human inventorship could potentially become more difficult.
In a recent UK High Court Decision [3], Mr Justice Smith ruled that an AI machine named “DABUS” could not be considered to be an “inventor” under the UK Patents Act 1977 because it is not a natural person and because, in any event, the right to obtain a patent application could not be transferred from an AI inventor to a patent applicant because an AI inventor would lack the necessary “legal personality” to own and transfer property rights. The European Patent Office came to a similar decision in respect of corresponding European patent applications filed naming DABUS as the sole inventor, albeit those cases are now under appeal. Thus the patent system in Europe generally does not (at least yet) recognise AI inventors.
However, there are still legitimate questions regarding who should rightly be named as the inventor in cases where an AI-entity does the “inventing”. Mr Justice Smith’s UK High Court decision positively left open the possibility that the owner of an AI machine which “invents” something could be named as the inventor (rather than the AI entity itself). This would potentially allow inventions made primarily by AI machines, where the AI machine owner has contributed minimally or not at all to the inventive distinction from the state of the art, to be patented by their owners in the UK in the future. Further development of the case law in this area (both in the UK and in Europe) is anticipated over the coming years, which will hopefully bring more certainty to patent applicants.
Conclusion
AlphaFold 2 demonstrates the game-changing scientific advancements that are achievable with AI, and we expect that such AI tools will open up new areas of technology for scientists and engineers to explore. Even as patent systems around the world hurriedly adapt to keep pace with technological developments that often outpace patent law, many AI-related inventions are patentable and patent applications in this field have been soaring globally. However, there are some questions remaining, and it will be interesting to see how scientists will use AI to research and generate new therapies.
[1] https://en.wikipedia.org/wiki/AlphaFold
[2] https://deepmind.com/blog/article/alphafold-a-solution-to-a-50-year-old-grand-challenge-in-biology
[3] https://www.bailii.org/ew/cases/EWHC/Patents/2020/2412.html
[4] https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2220018/
This article was prepared by HGF Senior Patent Directors Eleanor Purnell and Dr Andrew McGettrick.
























